Back in the spring of last year, a 45-year-old man went to the Brigham and Women's Hospital in Boston because of a coronavirus infection. Doctors treated him with steroids and discharged him five days later.
But the infection never went away — for 154 days. "He was readmitted to the hospital several times over the subsequent five months for recurrence of his COVID-19 infection and severe pneumonia," says infectious disease doctor Jonathan Li at Harvard Medical School, who helped treat the man.
"So this is an extraordinary individual," Li says.
So extraordinary in fact that this man's case is offering scientists surprising clues about where the new coronavirus variants emerged and why they're causing explosive outbreaks on three continents.
To be clear here, the man wasn't what doctors call a "long hauler," or a person who clears a coronavirus infection and then continues to have health problems for months. This man had living, growing virus in his body for five months, Li says. The same infection lasted for five months.
"That is one of the remarkable aspects of this case," Li says. "In fact, he was highly infectious even five months after the initial diagnosis."
This man had a severe autoimmune disease that required him to take drugs to suppress his immune system. So his body couldn't fight off the coronavirus infection as well as a healthy person could. He would get better for a while, and then the virus would counterattack. He would fall sick again. Eventually, he ended up in the intensive care unit. He passed away five months after the initial diagnosis.
Throughout the man's infection, Li and his colleagues ran an illuminating experiment. Every few weeks, the team extracted coronavirus from the man's body and sequenced the virus's genome.
Li couldn't believe what they found. "I was shocked," he says. "When I saw the virus sequences, I knew that we were dealing with something completely different and potentially very important."
The sequences showed Li and his team that the virus was changing very quickly inside the man's body. The virus wasn't picking up just one or two mutations at a time. But rather, it acquired a whole cluster of more than 20 mutations. Scientists had never seen SARS-CoV-2, the coronavirus that causes the disease COVID-19, mutate so quickly during the whole pandemic.
Furthermore, laboratory experiments have shown that some of those mutations help the virus bypass detection by antibodies.
"Toward the very end of his life, he was treated with monoclonal antibodies, from Regeneron," Li says. "And shortly thereafter, we saw evidence that suggested the virus was developing resistance or escaping from these antibodies as well."
Li and his colleagues published their findings in The New England Journal of Medicine in early November 2020 with little fanfare. Then about a month later, the pandemic took a surprising turn — and this peculiar case in Boston took on a new importance.
Scientists in the U.K. and South Africa announced they had detected new variants of the coronavirus. These variants were causing huge surges of COVID-19 in these countries.
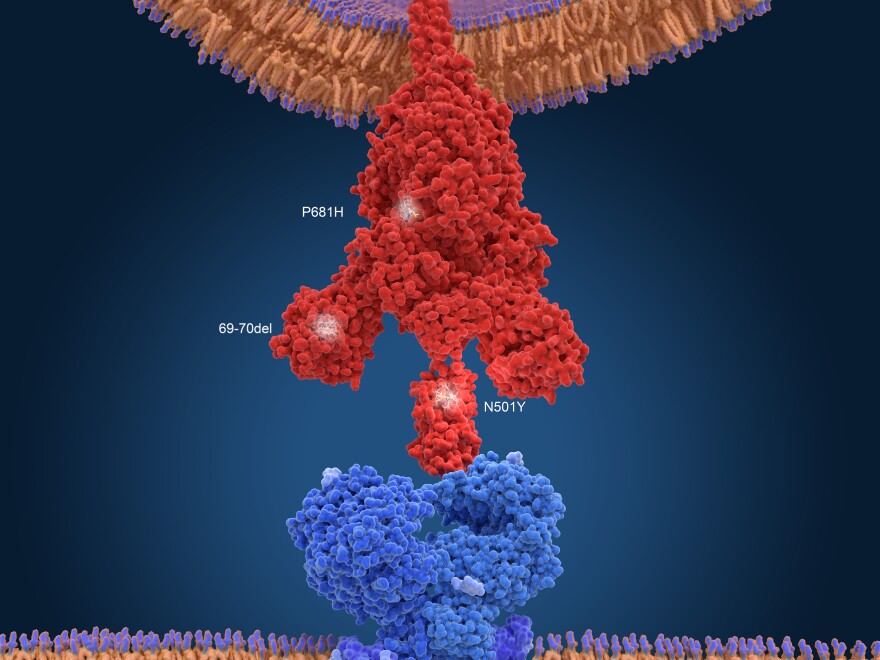
When researchers looked at the genes of these variants, guess what they found? A cluster of mutations that looked remarkably similar to the mutations found in the virus from the Boston patient. The sets of mutations weren't exactly identical, but they shared important characteristics. They both had about 20 mutations, and they shared several key ones, including a mutation (N501Y) known to help the virus bind more tightly to human cells and another mutation (E484K) known to help the virus evade antibody detection.
Since Li and his colleagues published their findings, several other teams have reported similar cases in which the virus evolved rapidly inside an immunocompromised person with a chronic coronavirus infection.
"So we have a number of examples, around the planet, in which patients' viruses suddenly have a whole mess of new mutations all at once," says virologist Jeremy Luban at the University of Massachusetts Medical School. And other cases have likely gone undetected, he says.
So scientists are starting to think the two phenomena could be related. That perhaps the new variants arose inside people similar to the man in Boston — that is, people who are immunocompromised and have long-term coronavirus infections.
"I think that's the leading theory," Luban says.
In other words, perhaps the coronavirus uses long-term infections as a mutational testing ground. While inside one person, the virus can try out all these different combinations of mutations and figure out, through trial and error, which ones are best at evading the immune system or helping the virus become more infectious.
Most of these viral versions probably don't spread beyond the chronically infected patient. But every once in a while, as the theory goes, a variant gets lucky, infects a large number of people and launches a new whole stage of the pandemic.
And this process is likely happening again right now, worldwide, in other immunocompromised patients. Eventually, these new variants could mutate again and create even more dangerous forms of the coronavirus.
Copyright 2021 NPR. To see more, visit https://www.npr.org.